Horibamultimodelpredictor: Difference between revisions
Jump to navigation
Jump to search
(Created page with "=Multi-Model Predictor= __TOC__ =Introduction= The Multi-Model Predictor is an Add-In component for Solo developed by HORIBA Scientific for use with HORIBA Fluorescence syste...") |
m (Scott moved page Horibamultimodelapply to Horibamultimodelpredictor: Horiba request name change.) |
||
| (3 intermediate revisions by the same user not shown) | |||
| Line 3: | Line 3: | ||
=Introduction= | =Introduction= | ||
The Multi-Model Predictor is an Add-In component for Solo developed by HORIBA Scientific for use with HORIBA Fluorescence systems. To enable the Add-In contact visit www.horiba.com/fluorescence. | The Multi-Model Predictor is an Add-In component for Solo developed by HORIBA Scientific for use with HORIBA Fluorescence systems. To enable the Add-In contact visit [https://www.horiba.com/fluorescence www.horiba.com/fluorescence]. | ||
=Reporting Interface= | =Reporting Interface= | ||
Reports can be generated using the following steps: | Reports can be generated using the following steps: | ||
* If it hasn't been specified already, add a default method file folder using the Edit>Options menu item. Do the same for a default log file location if needed. | |||
* Select a '''Method''' from the method drop-down menu. | * Select a '''Method''' from the method drop-down menu. | ||
* Select a log file from a folder containing both the log file and matching data files. | * Select a log file from a folder containing both the log file and matching data files. | ||
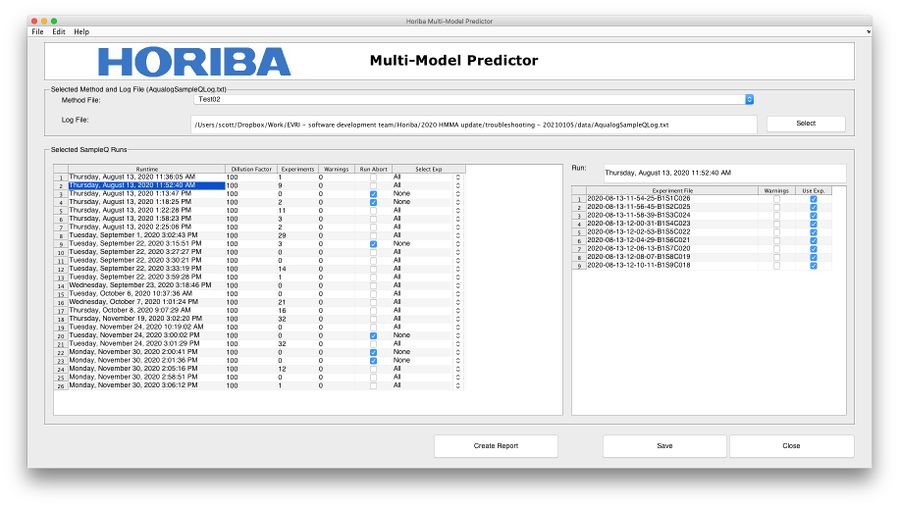
* View '''Runs''' in the run table. Experiments will be pre-selected based on ''abort'' condition. Use the '''Select Exp''' field to automatically adjust experiment selection. Note, right-clicking in the run table will show a context menu that allows for selecting all/no experiments for entire run table. | * View '''Runs''' in the run table. Experiments will be pre-selected based on ''abort'' condition. Use the '''Select Exp''' field to automatically adjust experiment selection. Note, right-clicking in the run table will show a context menu that allows for selecting all/no experiments for entire run table. | ||
* Further adjust experiment selection using the '''experiment table''' by manually checking the '''Use Exp.''' column. | * Further adjust experiment selection using the '''experiment table''' by manually checking the '''Use Exp.''' column. | ||
* Click the '''Create Report''' button to open a new window with a table of results. | |||
* Click '''Save''' from the new window so save a .csv file of the results. | |||
[[Image:HMMAReportInterface.jpg| 900px|HMMA Report]] | [[Image:HMMAReportInterface.jpg| 900px|HMMA Report]] | ||
=Method Interface= | |||
Use these steps to create and edit '''Method''' files: | |||
* From the '''File''' menu select '''Admin Login''' and enter user and password. | |||
* In the new panel the is shown, right-click and select '''New Method.''' | |||
* Name the method and, if desired, add a file name. | |||
* Drag and drop a [[Multiblocktool|Multi-Block]] model onto the new method. Note these can be models from within the Solo workspace or as saved .mat files. | |||
* Drag and drop models to apply. For classification models only a single model will be allowed. For regression models, multiple models can be used. Use right-click menu to remove models. | |||
* When a method has been assembled, use the right-click '''Validate Mehtod''' menu item to check that the sizes expected by the multiblock model are the same as those expected by the regression/classification models. | |||
* To remove a method, mover or delete the .met file from the parent folder. | |||
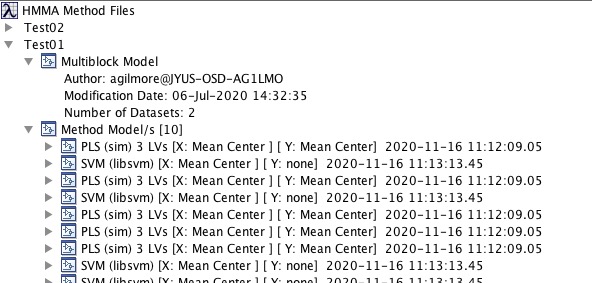
[[File:HMMAMethodEditTree.jpg| |HMMA Method Tree]] | |||
Latest revision as of 13:06, 22 March 2021
Multi-Model Predictor
Introduction
The Multi-Model Predictor is an Add-In component for Solo developed by HORIBA Scientific for use with HORIBA Fluorescence systems. To enable the Add-In contact visit www.horiba.com/fluorescence.
Reporting Interface
Reports can be generated using the following steps:
- If it hasn't been specified already, add a default method file folder using the Edit>Options menu item. Do the same for a default log file location if needed.
- Select a Method from the method drop-down menu.
- Select a log file from a folder containing both the log file and matching data files.
- View Runs in the run table. Experiments will be pre-selected based on abort condition. Use the Select Exp field to automatically adjust experiment selection. Note, right-clicking in the run table will show a context menu that allows for selecting all/no experiments for entire run table.
- Further adjust experiment selection using the experiment table by manually checking the Use Exp. column.
- Click the Create Report button to open a new window with a table of results.
- Click Save from the new window so save a .csv file of the results.
Method Interface
Use these steps to create and edit Method files:
- From the File menu select Admin Login and enter user and password.
- In the new panel the is shown, right-click and select New Method.
- Name the method and, if desired, add a file name.
- Drag and drop a Multi-Block model onto the new method. Note these can be models from within the Solo workspace or as saved .mat files.
- Drag and drop models to apply. For classification models only a single model will be allowed. For regression models, multiple models can be used. Use right-click menu to remove models.
- When a method has been assembled, use the right-click Validate Mehtod menu item to check that the sizes expected by the multiblock model are the same as those expected by the regression/classification models.
- To remove a method, mover or delete the .met file from the parent folder.